UPDATE 25/9/2017 I have a third biological replicate for L. crispatus growth that confirmed previous findings. I added a graph with average +/- standard deviations.
So, I have been growing the isolates that I ordered (see previous post) from DSM with mixed success.
-The bad news is that the Gardnerella vaginalis -80°C glycerol stock seems to be in bad shape. I inoculated twice successfully from this stock on NYCIII growth media (both liquid and agar plates), but after this it took multiple days for the culture to grow and lately they haven’t grown at all. I suspect this has to do with the aerobic condition in which they are stored. I will retry with plates and media that I will preincubate in anaerobic (N2+CO2) conditions, but there will be some influx of oxygen while inoculating, and at this point I am not sure that it is feasible to handle an study G. vaginalis outside of an anaerobic chamber. Other methods are still optional, such as using closed infusion flasks.
-The good news is that both Lactobacillus crispatus and Lactobacillus iners stocks seem to be growing well. I am using NYCIII medium for both, Lactobacillus crispatus grows within ~16 hours anaerobically, while Lactobacillus iners requires ~48 hours.
I present my first experiences with growing L. crispatus on glycogen.
Some background
The goal of this endeavor is to test whether vaginal bacteria can grow on glycogen, and are able to convert glycogen to lactic acid. Glycogen is an important carbohydrate present in the vagina, shown by old [1] and new research [2]. It is not yet understood how lactobacilli acidify the vagina of reproductive age women, and glycogen metabolism could be an important mechanism. Previously, Spear et al [3] have shown the presence of glycogen degrading enzymes in the vagina. The lactobacilli that were tested in this study showed no glycogen degrading capacity . Spear et al, asserts that the host excretes amylases (glycogen degrading enzymes) in order to assist acidification by Lactobacillus. However, the researchers did not use the most often encountered vaginal lactobacilli: Lactobacillus iners and Lactobacillus crispatus. Here I show initial growth experiments of L. crispatus on glycogen and amylase assay of the supernatants. I have gathered some evidence that Lactobacillus crispatus DSM 20584 strain can utilize glycogen as a source for growth.
Methods and results
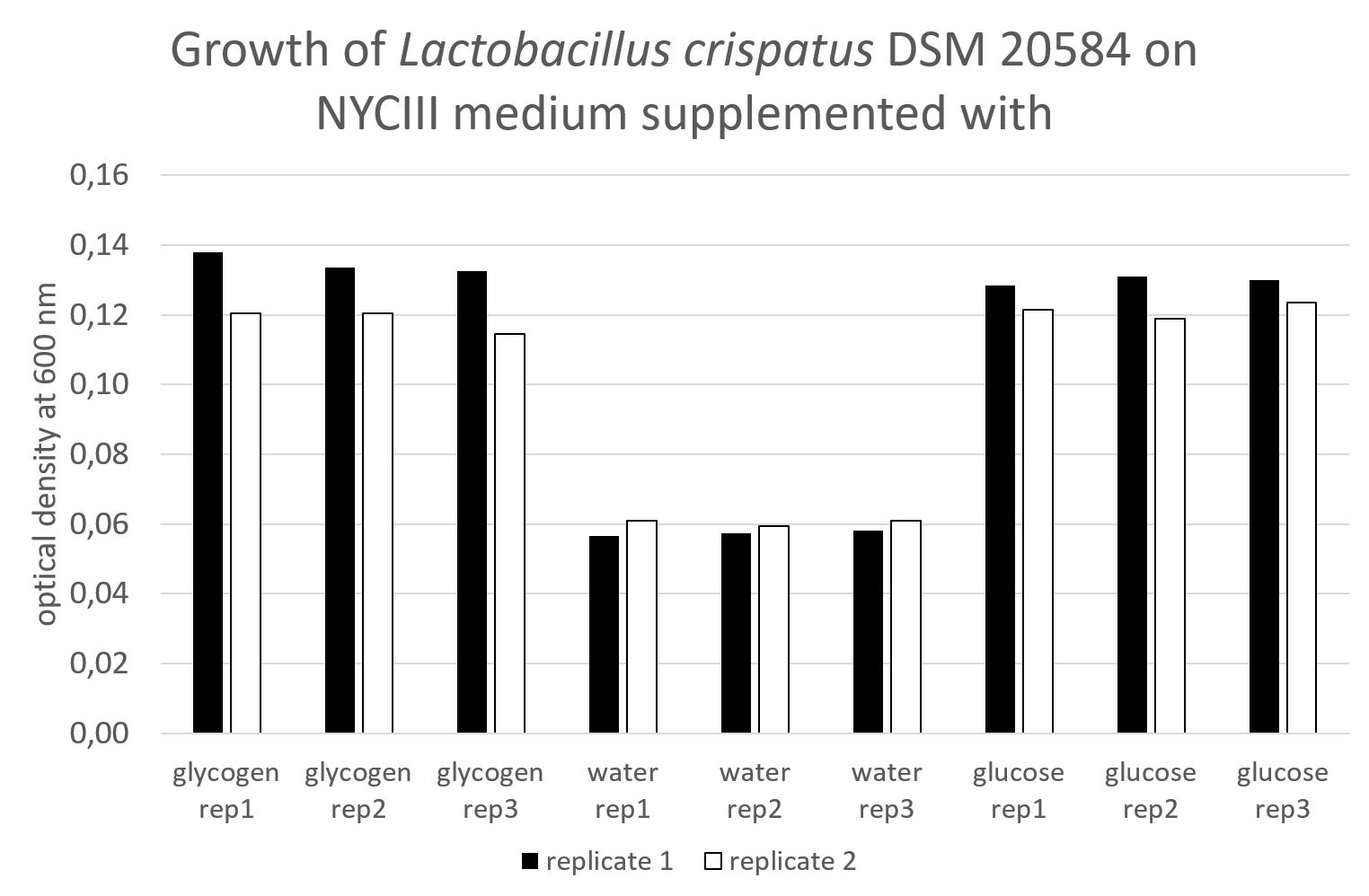
I have used NYCIII medium supplemented with either 0.5% glycogen or nothing (water, negative control) or 0.5% glucose as a positive control. To this end I have pipetted 100 uL of a 5% glucose solution or 5% glycogen solution or water and 900 uL of a 1.1x NYCIII medium where I have left out the glucose (see protocols). I have inoculated this with 100 uL of a preculture of L. crispatus DSM 20584 grown for >24 hours at 37°C anaerobically, without shaking. . For every condition, I used three wells as technical replicates. I also have empty controls without cells. Glycogen makes the solution a bit hazy so I wanted to make sure that any increase in optical density is not due to the glycogen itself but this was not the case. I have performed this experiment on two different occasions.
I mixed the culture by pipetting with a 1 mL pipet and diluted the culture 10x with water in a flatbottom 96-well plate to measure the cell density in the plate reader at OD 600. It looked like this

The data look like this: I’ll upload this to a platform once I have a bigger data file to share.
Updated graph with mean +/- standard deviations, unpaired t-test comparing OD600 on glycogen compared to water shows p-value of below .0005.



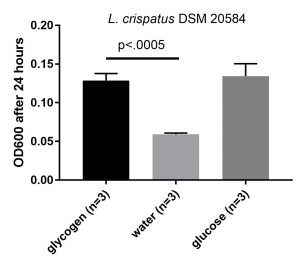
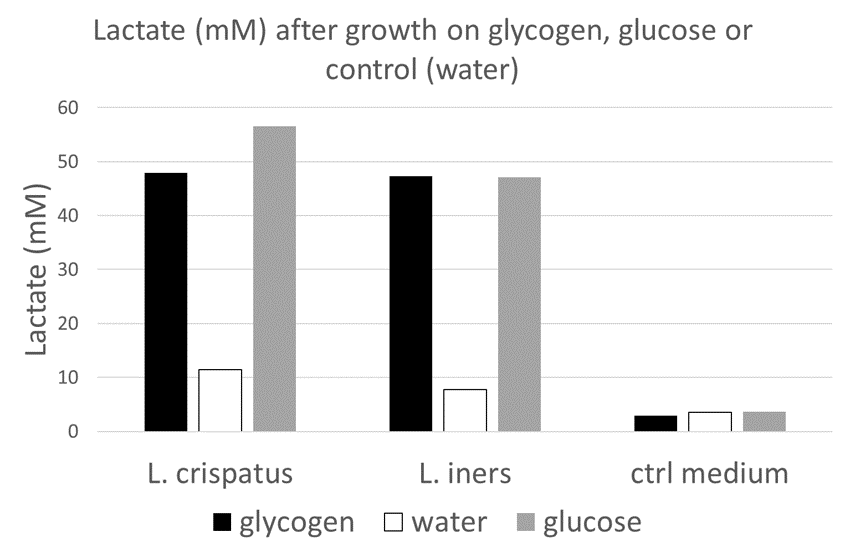
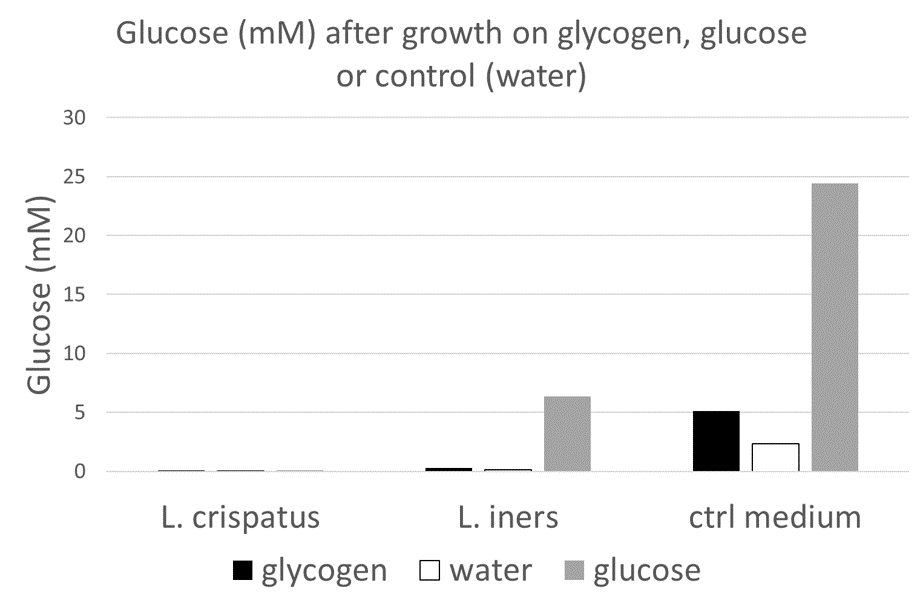
The optical density is increased when glycogen is added to the media, comparable to the increase by glucose. This is an indication that this particular strain can use glycogen for growth. Next, I wanted to see if I could detect the enzymes that L. crispatus uses to break down glycogen for uptake and metabolism.
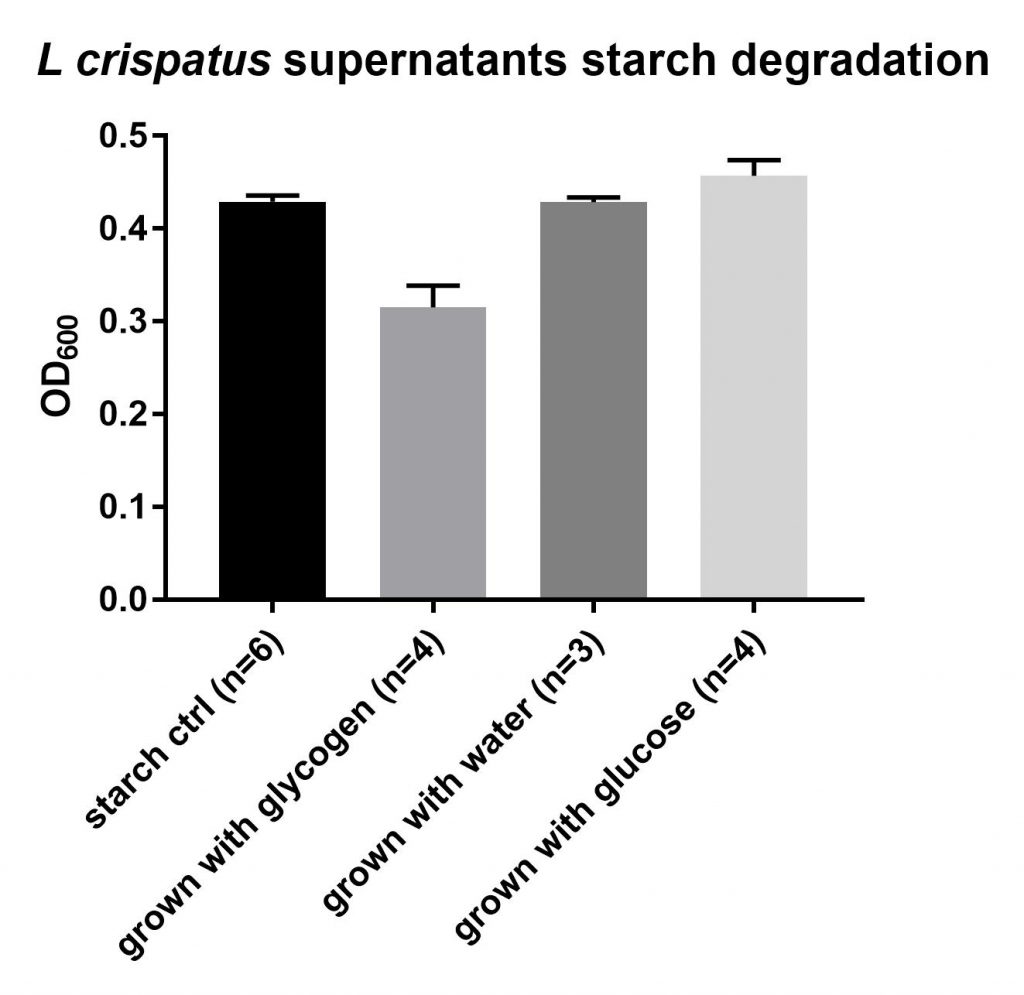
To this end, I spun down the cells for 20 minutes at 4°C and maximum speed (4754 rcf, 4499 rpm) and transferred the supernatants to plates I stored at -20°C. I wanted to test whether L. crispatus excreted any glycogen degrading enzymes in the supernatant. I added 50 uL of supernatant to 150 uL of a 1% starch solution in “amylase buffer” (100 mM Na-acetate+5mM CaCl2, pH 5.5). Starch is not the same as glycogen, the polymer has a different structure. The backbone of starch consists of the polymers amylose and amylopectin, which also consist mostly of glucose units but which are branched differently. Looking at Google images for these three polymers gives a good idea
I hope and expect that the enzymes that degrade most of the bonds between glucose units of glycogen will breakdown the bonds in amylose and amylopectin too. Starch has the advantage that it can be easily detected using iodine, an experiment that many of us already did in elementary school. I incubated this for ~24 hours at 37°C. I also checked after one and two hours but did not observe any breakdown of starch. I didn’t use any amylase control yet, only a reference calibration curve for starch. After incubation I added 10 uL, to 290 uL of iodine working solution and measured the absorption at 600 nm. I made a calibration curve using dilutions of the 1% starch concentration.
This experiment is not robust at all, since three times I measured three different things. So I will not share protocol and data yet. During the first measurement this calibration curve was not completely straight, not sure why, but the controls (starch with the supernatants of the empty control) showed an average of 7.6 gram/L starch, which is the concentration you expect. (150 uL of 10 gr/L starch + 50 uL of starchless liquid). However, in the wells with the L. crispatus supernatants something curious is going on. It seems that the amylase activity is found in the wells grown on glycogen whereas the wells grown with no carbon source (water) show the same concentration of starch as the controls (no breakdown). The glucose wells show some increase in starch. I am not sure how this happened, perhaps it has to do with pH, lactate presence, or even some growth in the wells. The wells did not look turbid, but growth can not be entirely ruled out.
The second time I did not freeze/thaw the supernatants but used fresh after spinning down the culture. I saw starch breakdown in the glycogen wells, indicating that L. crispatus excretes some soluble glycogen degrading enzyme in the supernatants (amylase?) whereas the H2O and glucose well did not show signs of starch breakdown. A third time I performed this experiment again with frozen sups I did not see any breakdown of starch.
So, I have to optimize this method, run controls with diluted amylase. I have to find out what’s going on with the increase in OD and finetune the protocol to get reproducible results. Perhaps it will turn out that this method is not robust enough, and I will have to resort to another way of measuring amylase activity.
On my long grocery wishlist with experiments:
- Improve amylase detection.
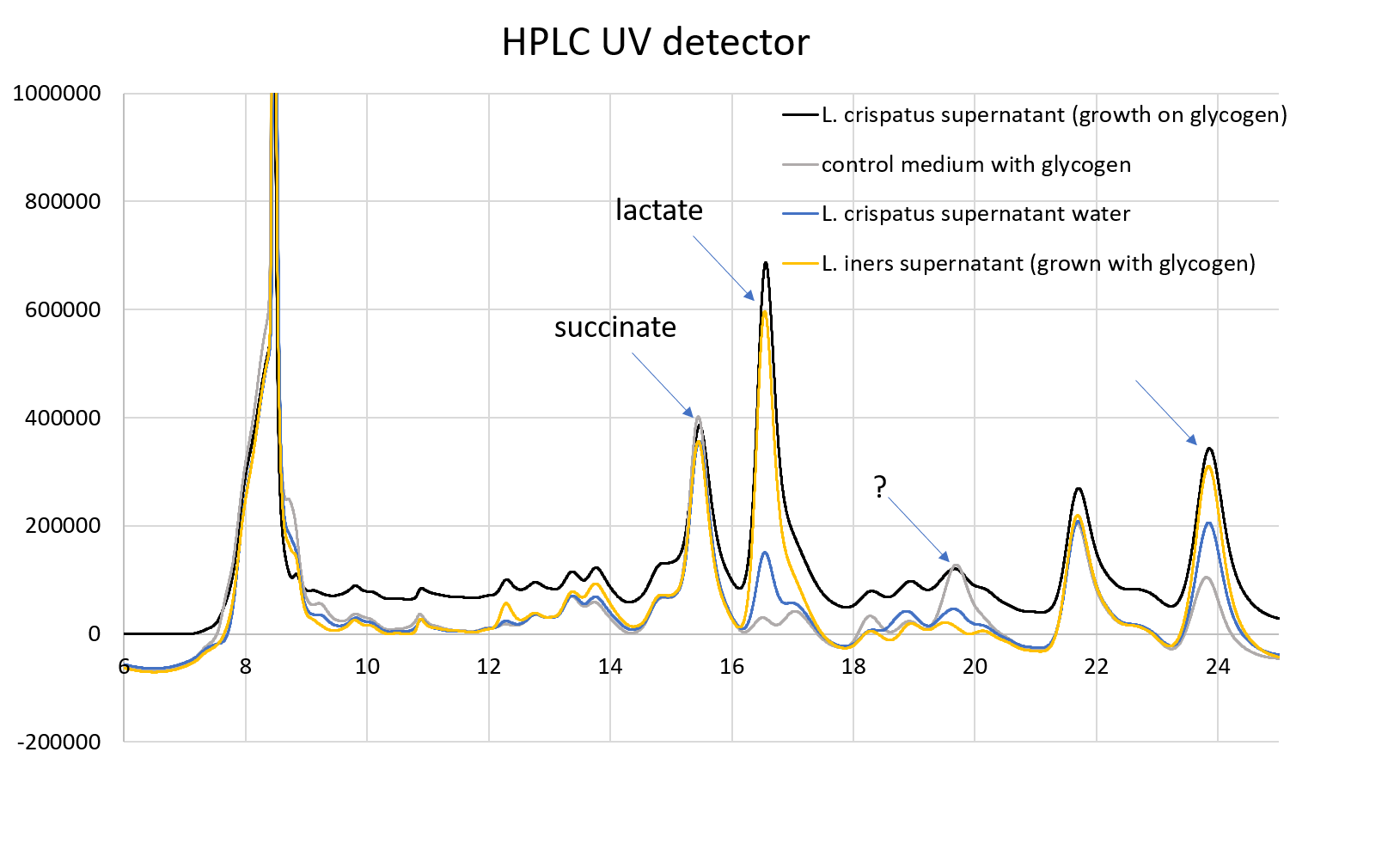
- Perform a HPLC on the supernatants to determine metabolite concentrations and gather more evidence of glycogen metabolism.
- Use HPLC to measure amylase products (glucose, maltose, trisaccharides etc).
Continue with Lactobacillus iners and Gardnerella vaginalis to study its glycogen degrading abilities.
- I would like to partially purify the amylases to perform proteomics, we have applied for a “Hotel”-grant at ZonMW to perform this analysis together with Winclove probiotics at Radboud UMC.
- Lastly, we want to extend the analysis to other strains of crispatus that were isolated by Remco Kort and his student Jorne Swanenburg in order to find the gene and enzyme involved in this amylase activity. Happy to announce that the biomedical/filosophy student Noa Fuks has offered her help to track down the amylase, while teaching herself some bioinformatics. Good luck Noa!
So, it’s a long grocery wishlist of experiments. If you have any comments, suggestions for alternative amylase activity assay, ideas, theories, criticism, please drop me a line.
1. Cruickshank, R., The conversion of the glycogen of the vagina into lactic acid. Journal of Pathology and Bacteriology, 1934. 29.
2. Mirmonsef, P., et al., Free glycogen in vaginal fluids is associated with Lactobacillus colonization and low vaginal pH. PloS one, 2014. 9(7): p. e102467.
3. Spear, G.T., et al., Human alpha-amylase present in lower-genital-tract mucosal fluid processes glycogen to support vaginal colonization by Lactobacillus. The Journal of infectious diseases, 2014. 210(7): p. 1019-1028.